Infection mechanisms of bacteriophages targeting motile bacteria
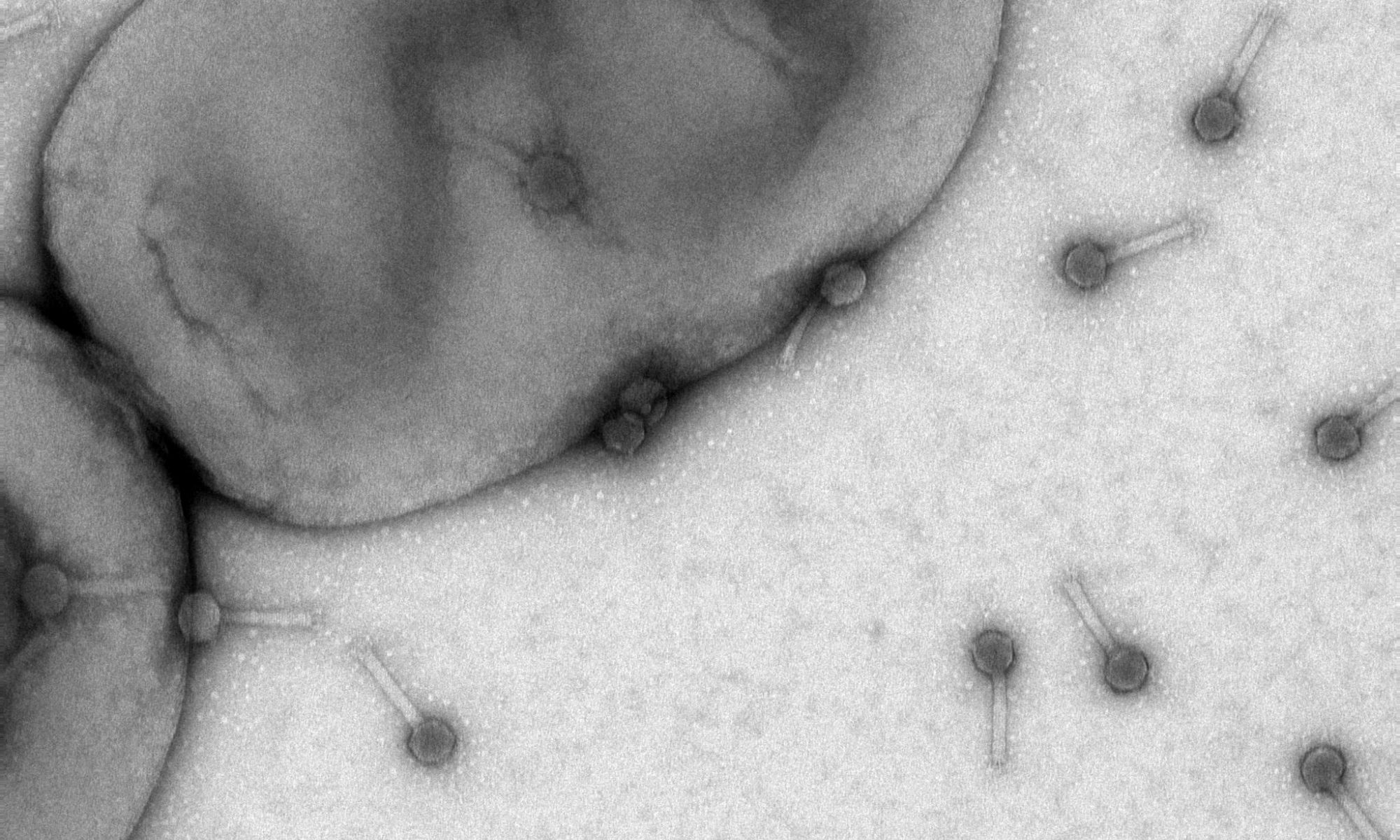
Bacteriophages are viruses of bacteria that use the host metabolism for propagation of new phage particles. One unique group of bacteriophages, named flagellotropic phages, are targeting the bacterial flagellum and rotation of the flagellar filament is an essential requirement for infection. Flagellar-driven motility and chemotaxis enables motile bacteria to move away from harmful and towards beneficial chemicals.
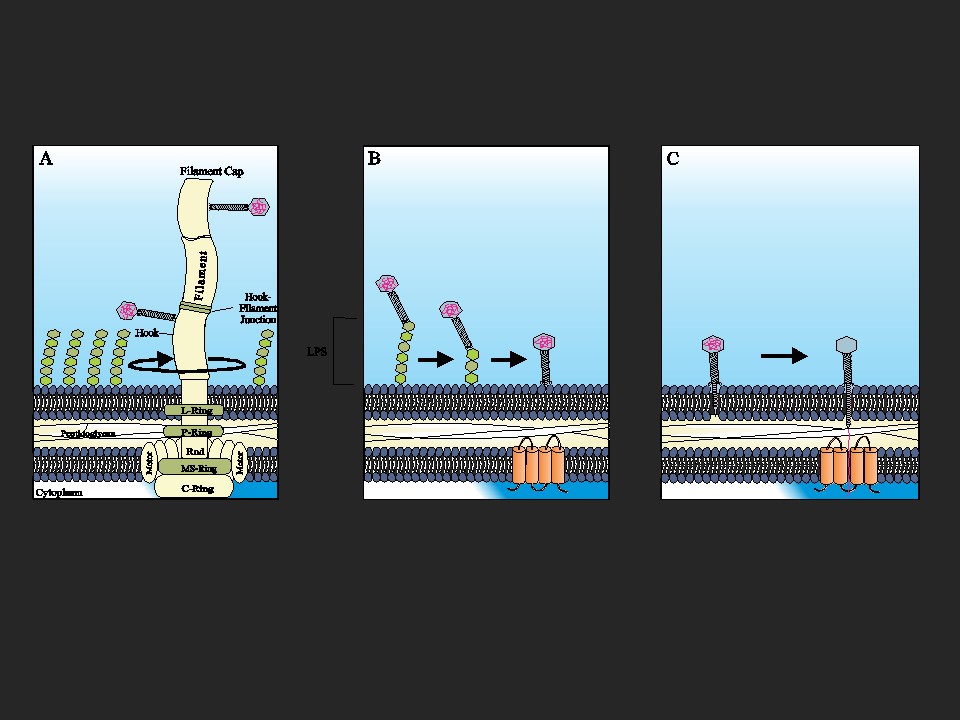
The soil bacterium Agrobacterium sp. H13-3 is the specific host for the flagellotropic phage 7-7-1. Co-evolution of bacteria and their phages resulted in the development of species-specific phage-host interactions. Previous research by the Scharf lab discovered that the flagellum and lipopolysaccharides (LPS) of A. sp. H13-3 serve as essential primary and secondary surface receptors for phage 7-7-1. However, nothing is known about the mechanisms of phage adsorption to the flagellum, translocation along the flagellum, or the subsequent infection steps. The overarching goal of this project is to characterize the specific processes involved in flagellotropic phage infection and host adaptation.
Salmonella enterica serovar Typhimurium is infected by flagellotropic phage Chi. Infection requires CCW flagellar rotation, however, details about the infection mechanism are unknown. The goal of this specific part of the project is identifying secondary cell surface receptors that are essential for the infection of S. Typhimurium by Chi phage using a high-throughput screening method of a barcoded S. Typhimurium 14028 gene-knockout library.
References
R.R. Sonani, Esteves N.C., Scharf B.E.*, Egelman E.H.* (2024) “Cryo-EM structure of flagellotropic bacteriophage Chi” Structure, 024 Apr 3:S0969-2126(24)00093-5. doi: 10.1016/j.str.2024.03.011. Online ahead of print. * Co-corresponding authors.
R.R. Sonani, Esteves N.C., Horton A.A., Kelly R.J., Sebastian A.L., Wang F., Kreutzberger M.A.B., Leiman P.G.*, Scharf B.E.*, Egelman E.H.* (2023) “Neck and capsid architecture of the robust Agrobacterium phage Milano” Springer Nature Communications Biology; 6(1):921. * Co-corresponding authors.
R.R. Sonani, Palmer L.K., Esteves N.C., Horton A.A., Sebastian A.L., Kelly R.J., Wang F., Kreutzberger M.A.B., Russell W.K., Petr G. Leiman P.G.*, Scharf B.E.*, Egelman E.H.* (2024) “An extensive disulfide bond network prevents tail contraction in Agrobacterium tumefaciens phage Milano” Springer Nature Communications Biology; 15(1):756. * Co-corresponding authors.
N.C. Esteves, Bigham D.N., Scharf B.E. (2023) Phages on filaments: A genetic screen elucidates the complex interactions between Salmonella enterica flagellin and bacteriophage Chi, PLOS Pathogens; 19(8).
N.C. Esteves & Scharf B.E. (2022) Flagellotropic bacteriophages: opportunities and challenges for antimicrobial applications. International Journal of Molecular Sciences 23(13):7084, 1-23.
F. Gonzalez & Scharf B.E. (2021) Identification of receptor binding proteins in flagellotropic Agrobacterium Phage 7-7-1. Viruses, 13(7), 1-21. (DOI: 10.3390/v13071267)
N.C. Esteves, Porwollik S., McClelland M., Scharf B.E. (2021) The multi-drug efflux system AcrABZ-TolC is essential for infection of Salmonella Typhimurium by the flagellum-dependent bacteriophage Chi. Journal of Virology, 59(11), 1-13. (DOI: 10.1128/JVI.00394-21)
X. Li, Gonzalez F., Esteves N., Scharf B.E., Chen J. (2020) Formation of phage lysis patterns and implications on co-propagation of phages and motile host bacteria. PLoS Computational Biology; 16(3), 1-22. (DOI: 10.1371/journal.pcbi.1007236)
D. William, Schleupner B., Varano A.C, Alden N., Gonzalez F., Casasanta M., Scharf B.E., Dukes M., Kelly D. (2019) Liquid-cell electron tomography of biological systems. Nano Letters, 19, 6734-6741. (DOI: 10.1021/acs.nanolett.9b01309)
F. Gonzalez, Helm R.F., Broadway K.M, Scharf B.E. (2018) More than rotating flagella: LPS as a secondary receptor for flagellotropic phage 7-7-1; Journal of Bacteriology, 200(19). (DOI: 10.1128/JB.00363-18)
J.Y. Yen, Broadway K., and Scharf B. E. (2012) Minimum requirements of flagellation and motility for the infection of Agrobacterium sp. H13-3 by the flagellotropic bacteriophage 7-7-1. Applied & Environmental Microbiology 78, 7216-7222. (DOI:10.1128/AEM.01082-12)
D. Wibberg D, Blom J., Jaenicke S., Kollin F., Rupp O., Scharf B., Schneiker-Bekel S., Sczcepanowski R., Goesmann A., Setubal J.C., Schmitt R., Pühler A., Schlüter A. (2011) Complete genome sequencing of Agrobacterium sp. H13-3, the former Rhizobium lupini H13-3, reveals a tripartite genome consisting of a circular and a linear chromosome and an accessory plasmid but lacking a tumor-inducing Ti-plasmid. Journal of Biotechnology, 50-62.(DOI:10.1016/j.jbiotec.2011.01.010)
B. Scharf (2002) Real-time imaging of fluorescent flagellar filaments of Rhizobium lupini H13-3: Flagellar rotation and pH-induced polymorphic transitions. Journal of Bacteriology, 5979-5986. (PMCID:PMC135403 )
B. Scharf, Schuster-Wolff-Bühring H., Rachel R., Schmitt R. (2001) Mutational analysis of the Rhizobium lupini H13-3 and Sinorhizobium meliloti flagellin genes: Importance of flagellin A for flagellar filament structure and transcriptional regulation. Journal of Bacteriology, 5334-5342. (PMCID:PMC95416)